In this article, we introduce how to use “Molecular Nodes”, an add-on that allows you to handle and visualize molecular data in Blender.
Molecular Nodes allows you to import, animate, and manipulate molecular data in Geometry Nodes.
The Molecular Nodes add-on works only with Blender 3.2 or higher, so this article uses Blender 3.3LTS.
This add-on requires Python operations to use some functions, but it is not difficult, so we encourage you to try it.
How to Install
Installing Addon
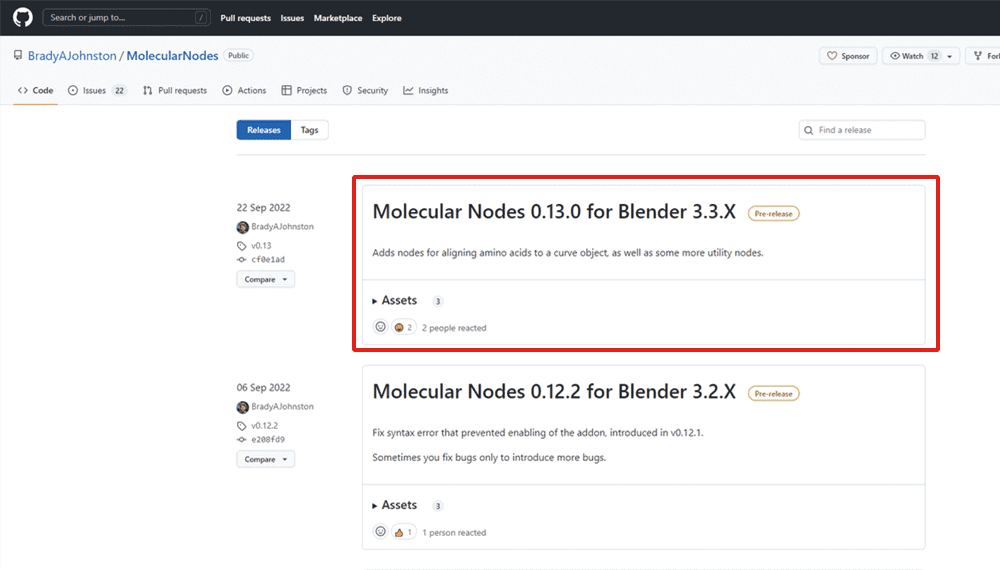
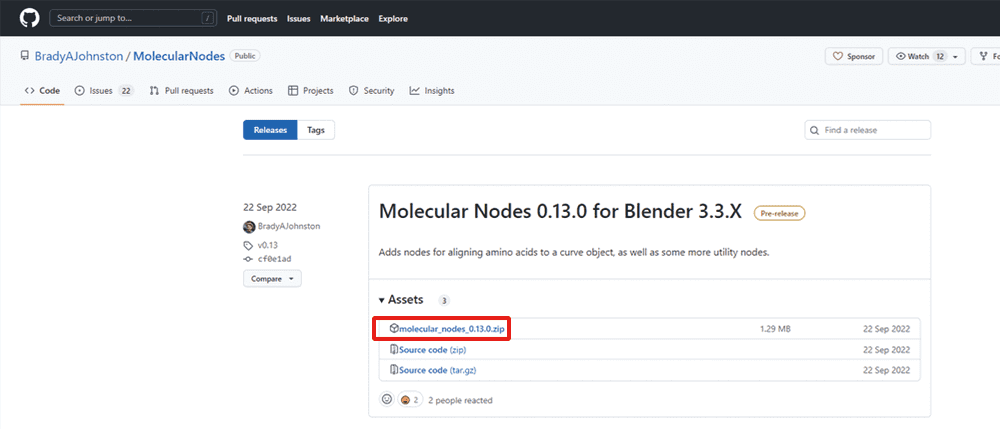
Go to GitHub at the following URL and download the zip file of the latest release:
Save the downloaded zip file in any location without unzipping.
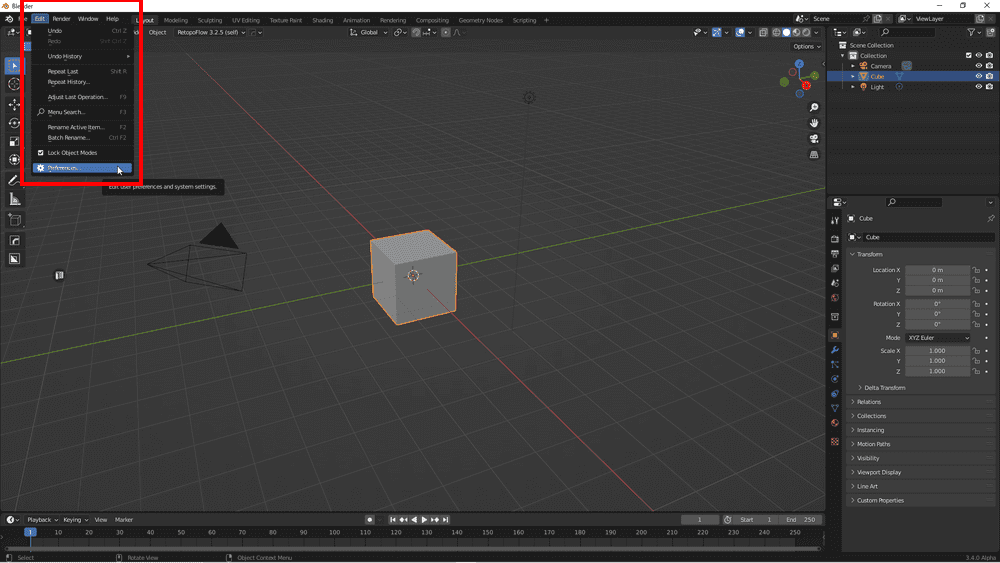
Start Blender, go to Edit→Preferences and select Add-ons.
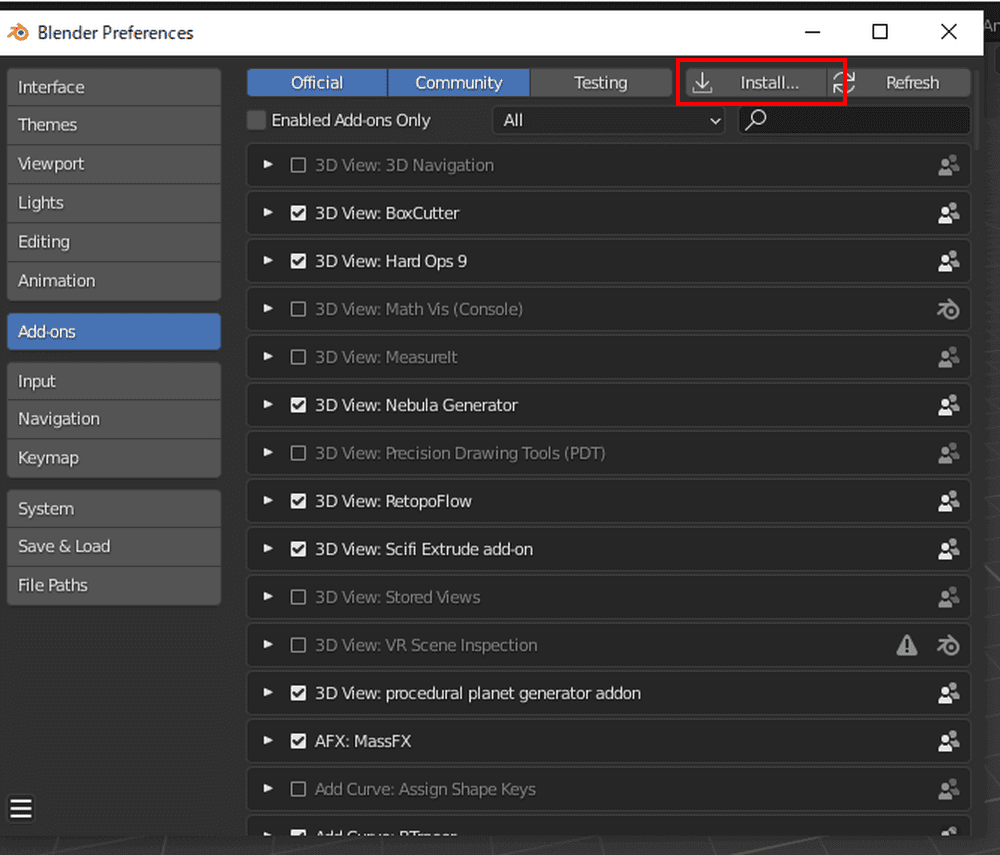
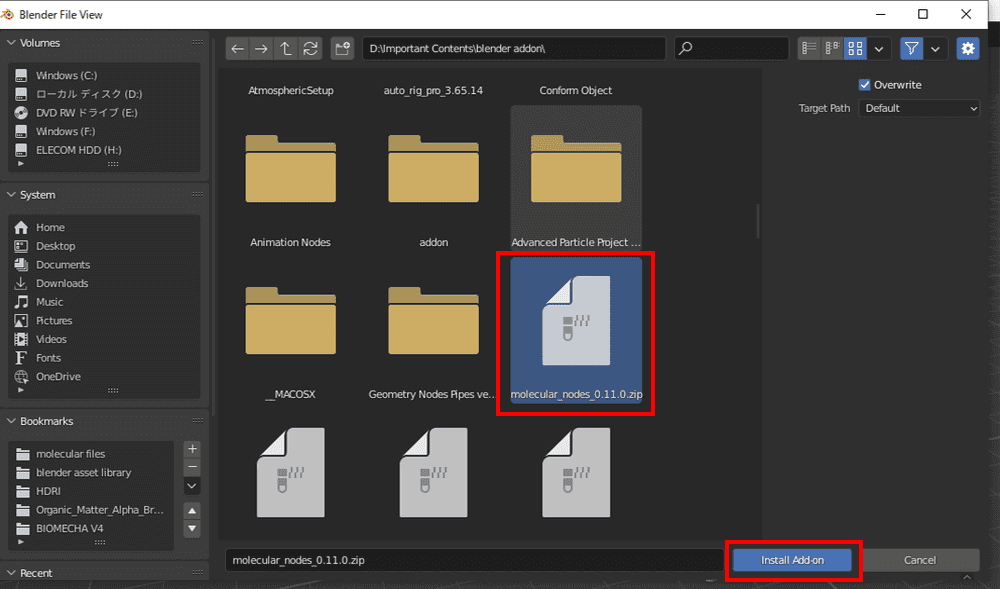
Click Install in the upper right corner. Select the zip file you just saved and install.
When the add-on appears, check the box next to the name to activate it.
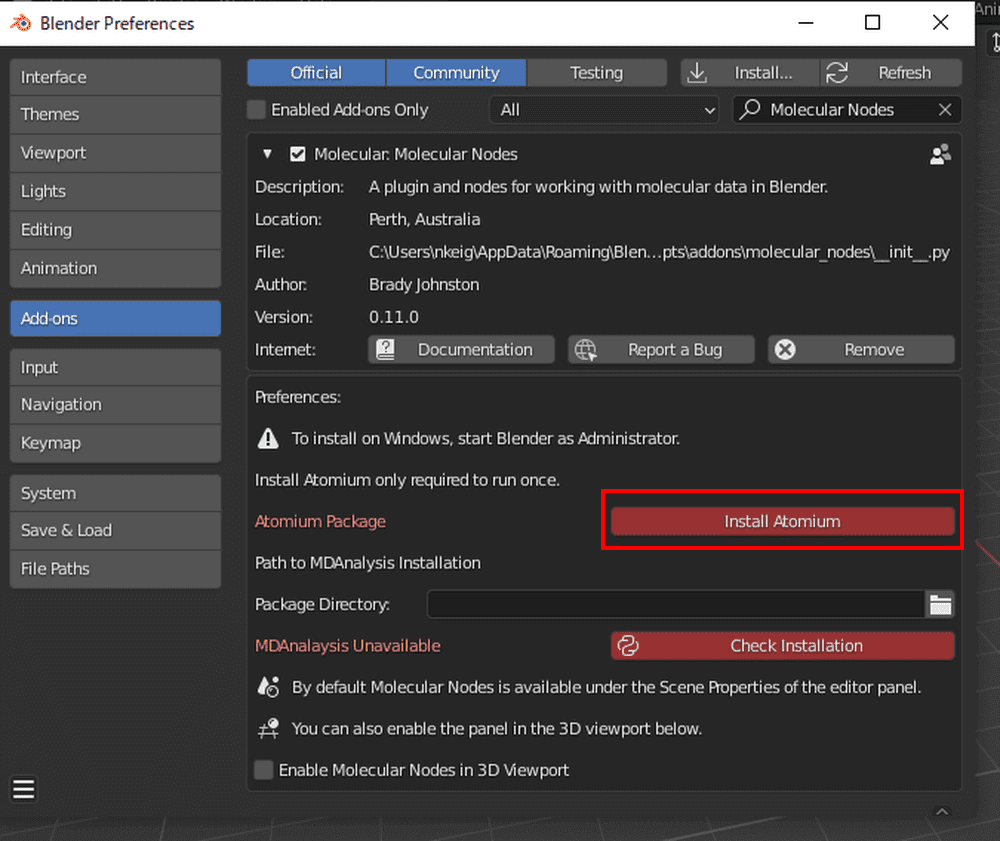
Click the triangle next to the checkbox to display the details.
Click on “Install Atomium”, which is shown in red.
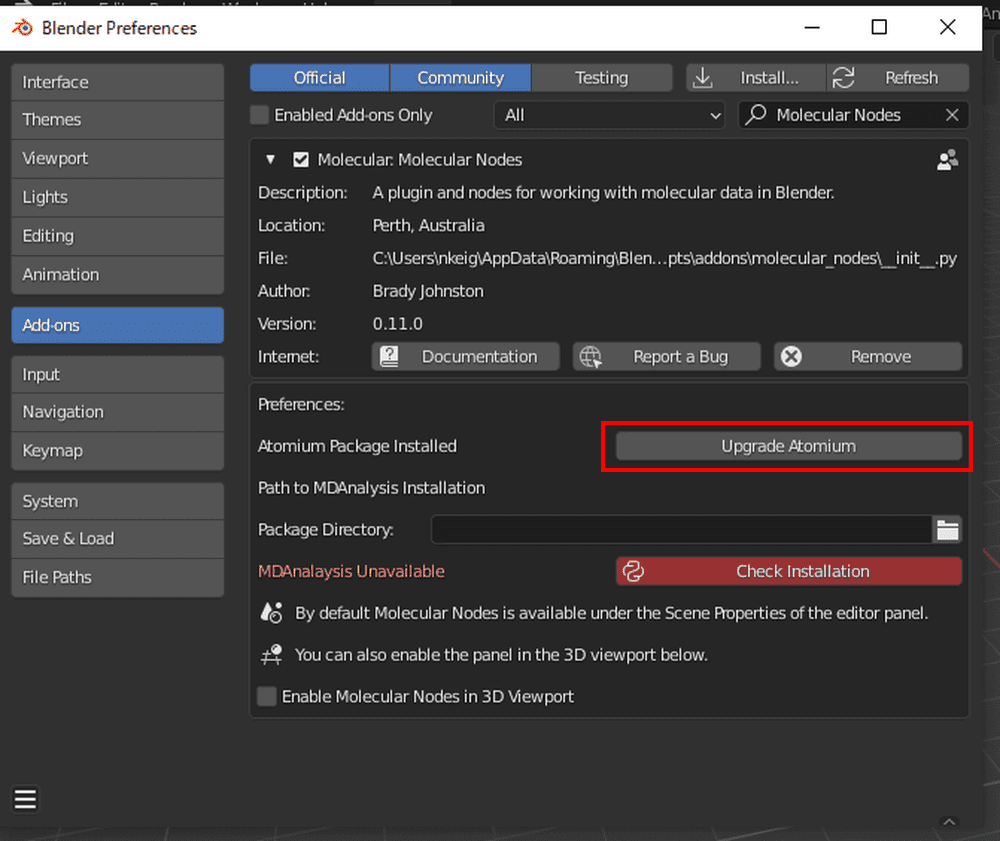
Wait for a while: when the color changes to gray, the installation of Atomium is complete.
Installing MDAnalysis
To read complex molecular data or data with special extensions, install MDAnalysis and associate it with the Molecular Nodes add-on.
If you only want to use some of the add-on’s features, you can skip installing MDAnalysis; we suggest you try it only if you can afford to do so.
Blender 3.1 or higher is bundled with Python 3.10 or higher, so install Python 3.10 or higher.
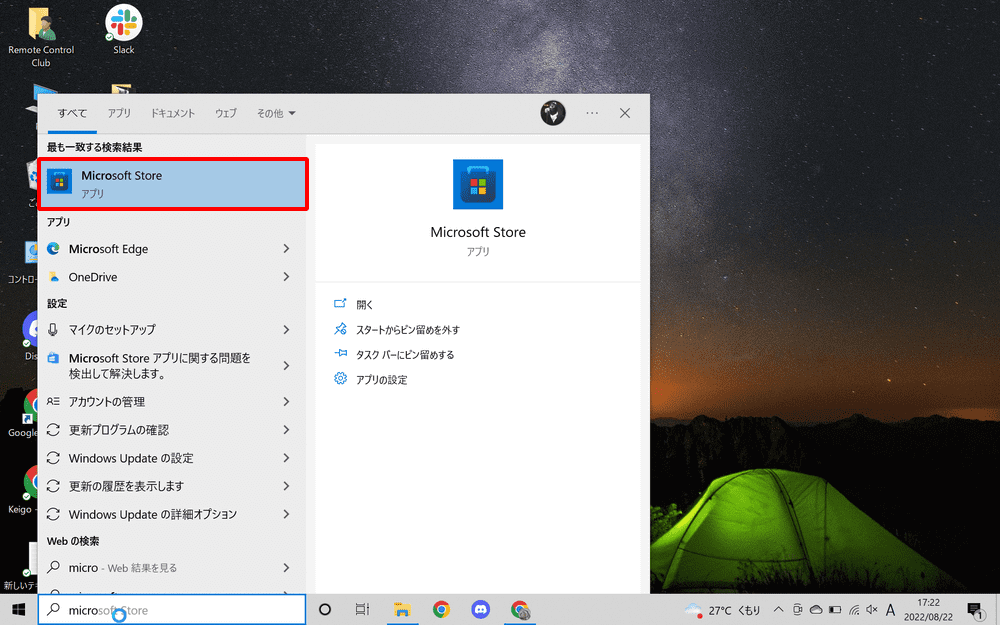
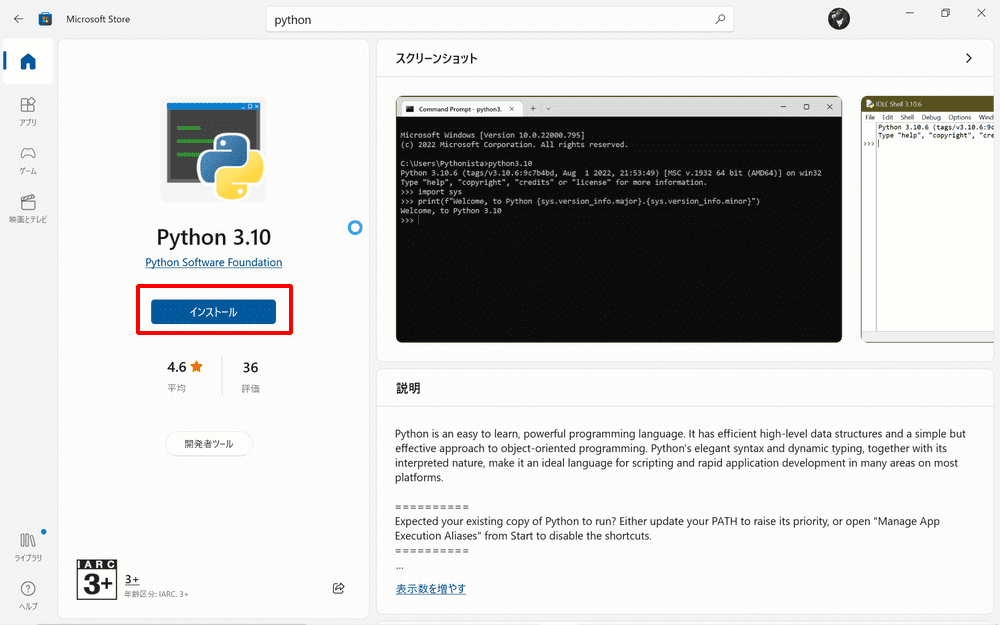
Type “Microsoft Store” in the search field of the taskbar and launch it.
Search for Python in Microsoft Store and install Python 3.10.
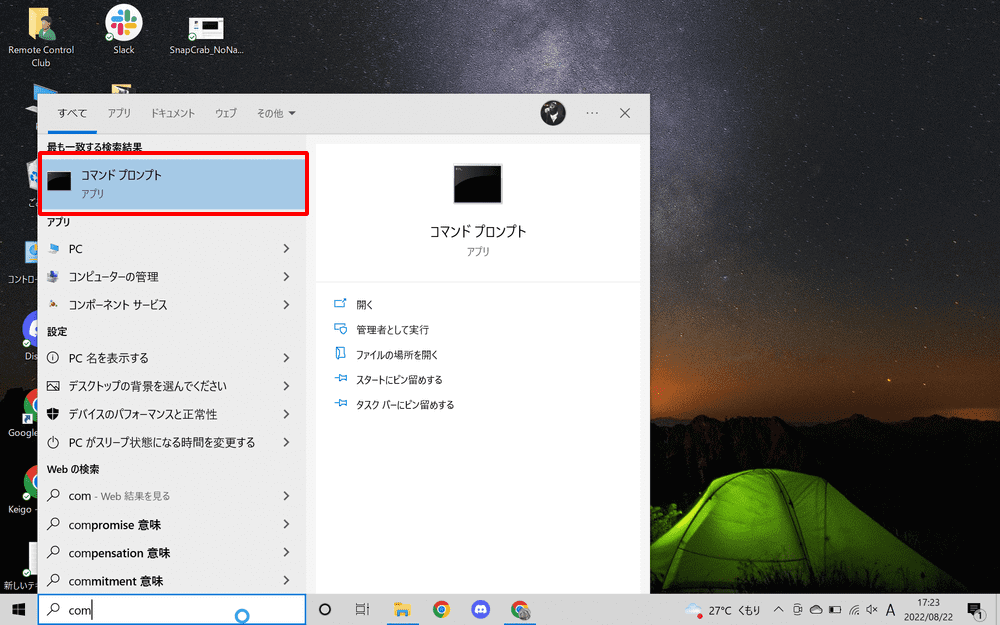
Once Python has been successfully installed, open the Command Prompt.
Type “cmd” or “command prompt” in the search field of the taskbar to start it.
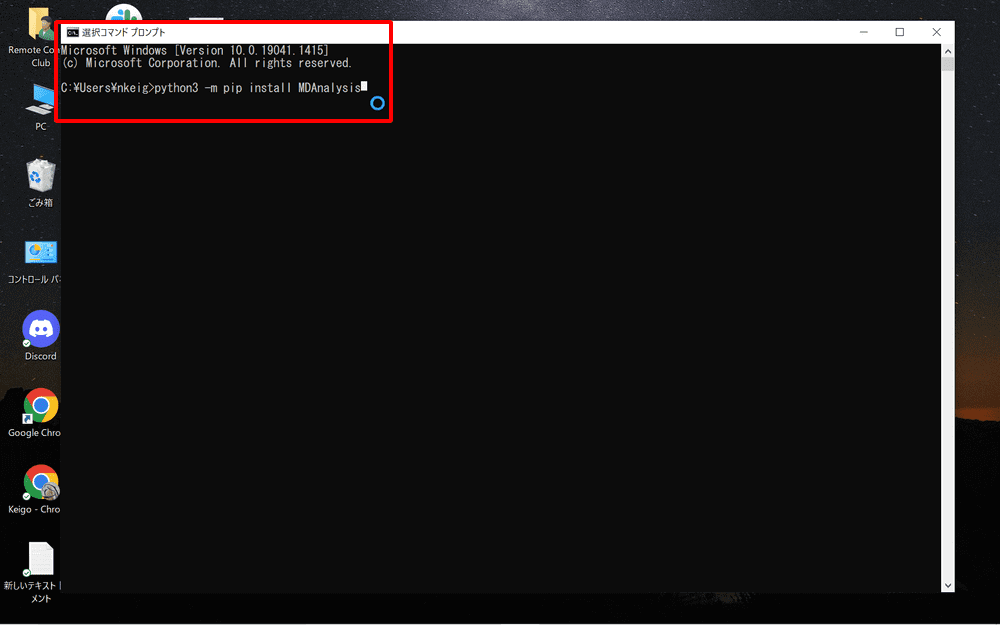
After launching the command prompt, copy and paste the following command:
python3 -m pip install MDAnalysis
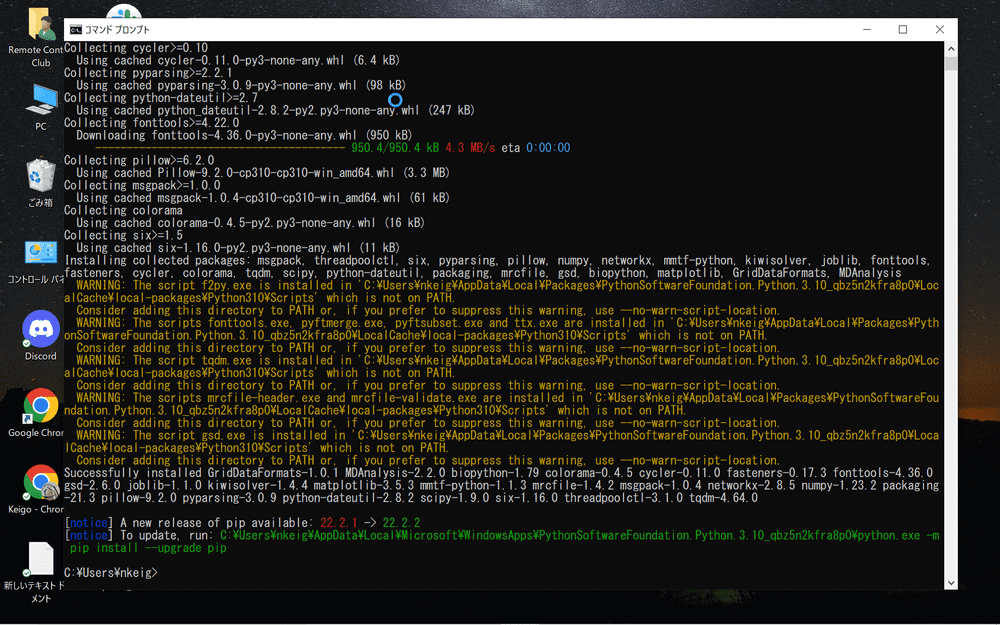
MDAnalysis will be installed.
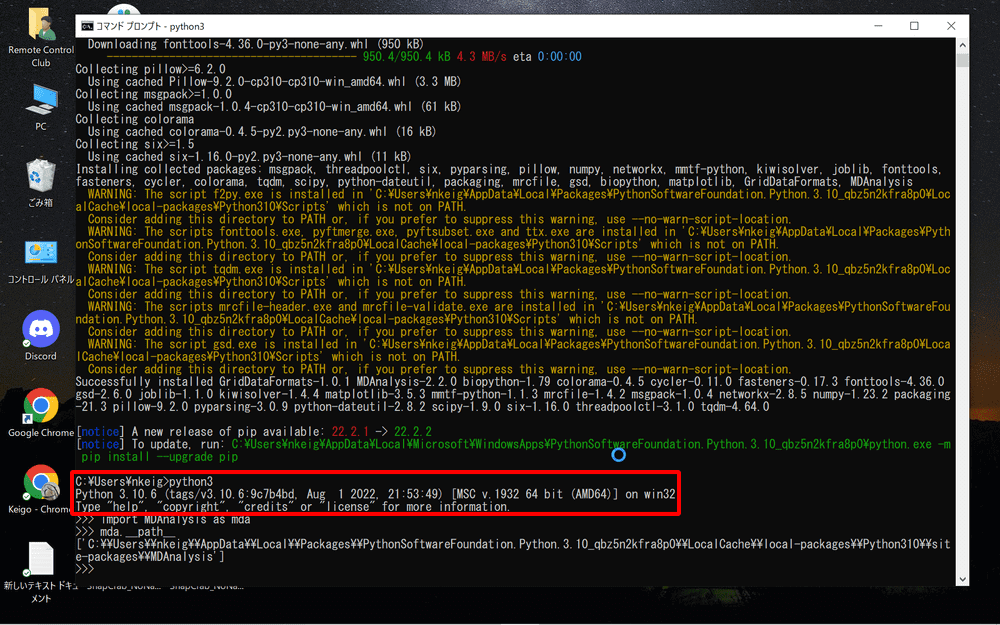
Once the installation is complete, enter “python3” to start the Python command line.
Next, enter the following command to display the MDAnalysys path:
import MDAnalysis as mda
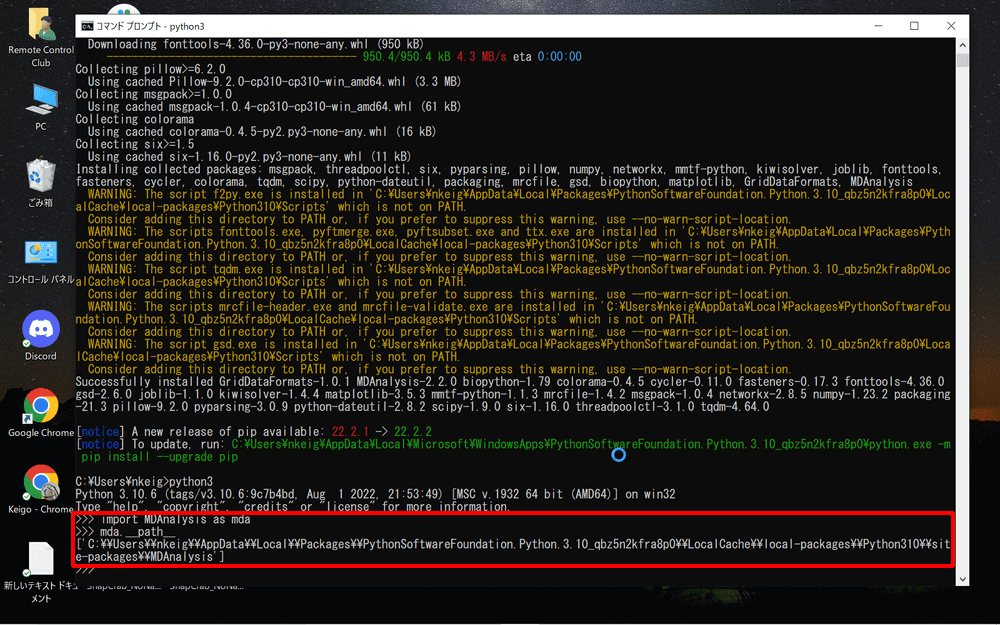
mda.__path__
Copy the path starting with C: and return to Blender.
For my case in the example, the path will be “C:\\Users\\nkeig\\AppData\\Local\\Programs\\Python\\Python310\\lib\\site-packages”
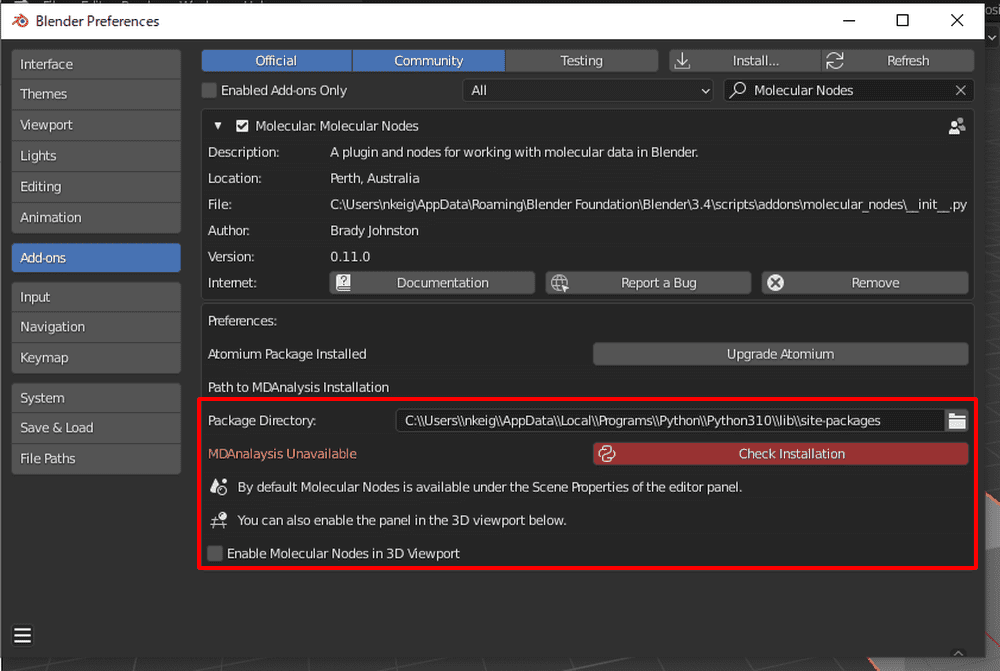
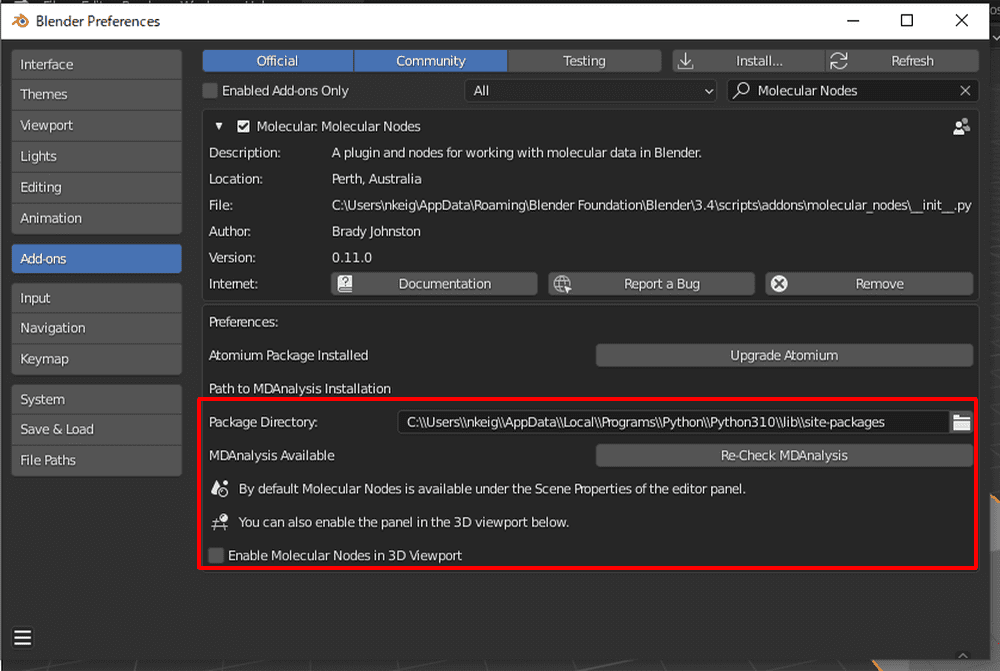
After returning to Blender, paste the path into the Molecular Nodes add-on’s Package Directory and click Check Installation.
It is successful when the color turns gray (after a while).
This completes all installation of Molecular Nodes.
The specific usage of the Molecular Nodes Addon is explained in detail in the following article:
For questions about STYLY, bug reports, and improvement requests, please contact the STYLY FORUM
https://en.forum.styly.cc/support/discussions
Certified (QA) by Shota Shawn Yoshizawa
Edited by SASAnishiki
Translated by passerby1
![[Blender] How to Handle Molecular Data with “Molecular Nodes” Addon (Part 2)](https://styly.cc/wp-content/uploads/2022/08/2-30-160x160.png)